Bromine »
PDB 2xpn-3bnq »
3bnq »
Bromine in PDB 3bnq: Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)
Protein crystallography data
The structure of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative), PDB code: 3bnq
was solved by
J.Kondo,
E.Westhof,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 49.79 / 2.00 |
| Space group | C 1 2 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 71.266, 76.056, 55.912, 90.00, 117.06, 90.00 |
| R / Rfree (%) | 22.9 / 26.9 |
Other elements in 3bnq:
The structure of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative) also contains other interesting chemical elements:
| Strontium | (Sr) | 8 atoms |
| Potassium | (K) | 2 atoms |
Bromine Binding Sites:
The binding sites of Bromine atom in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)
(pdb code 3bnq). This binding sites where shown within
5.0 Angstroms radius around Bromine atom.
In total 4 binding sites of Bromine where determined in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative), PDB code: 3bnq:
Jump to Bromine binding site number: 1; 2; 3; 4;
In total 4 binding sites of Bromine where determined in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative), PDB code: 3bnq:
Jump to Bromine binding site number: 1; 2; 3; 4;
Bromine binding site 1 out of 4 in 3bnq
Go back to
Bromine binding site 1 out
of 4 in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)
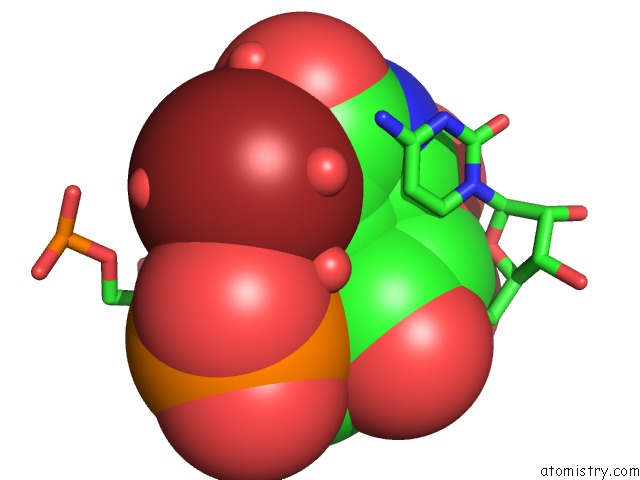
Mono view
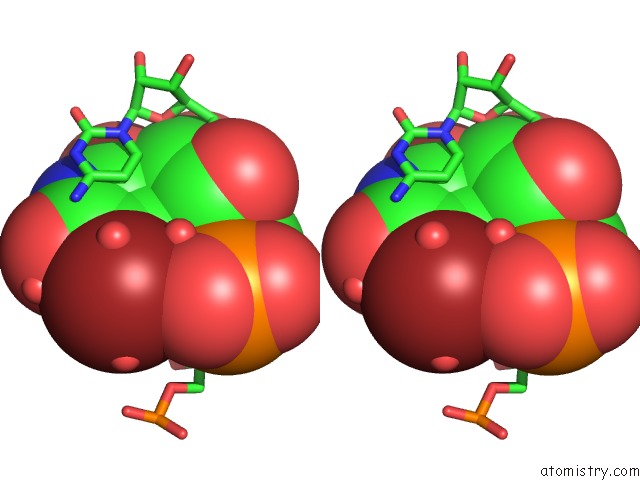
Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 1 of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative) within 5.0Å range:
|
Bromine binding site 2 out of 4 in 3bnq
Go back to
Bromine binding site 2 out
of 4 in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)

Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 2 of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative) within 5.0Å range:
|
Bromine binding site 3 out of 4 in 3bnq
Go back to
Bromine binding site 3 out
of 4 in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)
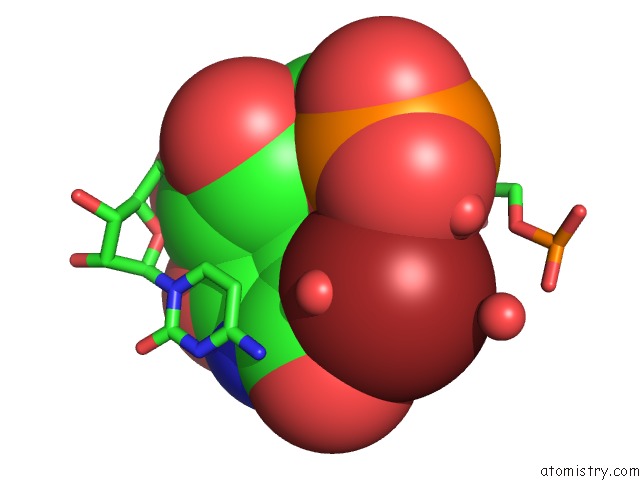
Mono view

Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 3 of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative) within 5.0Å range:
|
Bromine binding site 4 out of 4 in 3bnq
Go back to
Bromine binding site 4 out
of 4 in the Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative)

Mono view
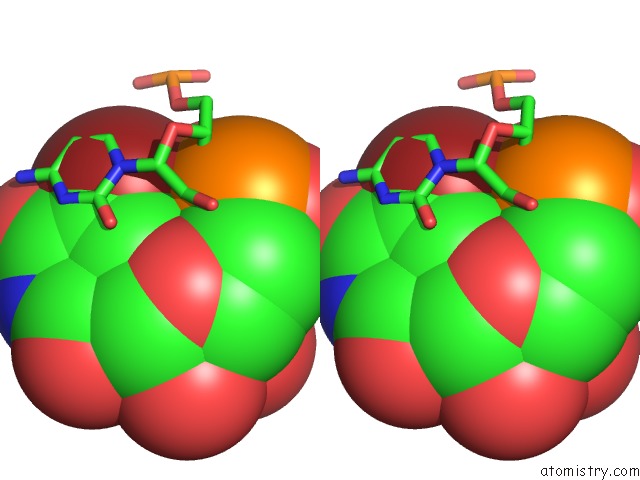
Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 4 of Crystal Structure of the Homo Sapiens Mitochondrial Ribosomal Decoding Site in the Presence of SRCL2 (A1555G Mutant, Br-Derivative) within 5.0Å range:
|
Reference:
J.Kondo,
E.Westhof.
The Bacterial and Mitochondrial Ribosomal A-Site Molecular Switches Possess Different Conformational Substates Nucleic Acids Res. V. 36 2654 2008.
ISSN: ISSN 0305-1048
PubMed: 18346970
DOI: 10.1093/NAR/GKN112
Page generated: Mon Jul 7 05:03:31 2025
ISSN: ISSN 0305-1048
PubMed: 18346970
DOI: 10.1093/NAR/GKN112
Last articles
F in 4IVWF in 4IVY
F in 4IVO
F in 4IV4
F in 4IVM
F in 4IV2
F in 4IUI
F in 4IN4
F in 4IU7
F in 4ITI