Bromine »
PDB 5t7a-5v1k »
5tmm »
Bromine in PDB 5tmm: Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid
Protein crystallography data
The structure of Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid, PDB code: 5tmm
was solved by
J.C.Nwachukwu,
S.Srinivasan,
N.E.Bruno,
J.Nowak,
D.J.Kojetin,
O.Elemento,
J.A.Katzenellenbogen,
K.W.Nettles,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 45.64 / 2.20 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 55.090, 82.520, 58.740, 90.00, 111.14, 90.00 |
| R / Rfree (%) | 20.5 / 25.1 |
Bromine Binding Sites:
The binding sites of Bromine atom in the Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid
(pdb code 5tmm). This binding sites where shown within
5.0 Angstroms radius around Bromine atom.
In total only one binding site of Bromine was determined in the Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid, PDB code: 5tmm:
In total only one binding site of Bromine was determined in the Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid, PDB code: 5tmm:
Bromine binding site 1 out of 1 in 5tmm
Go back to
Bromine binding site 1 out
of 1 in the Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid
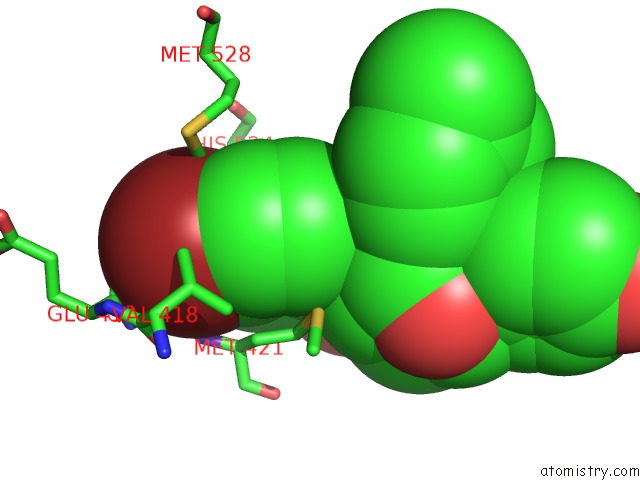
Mono view
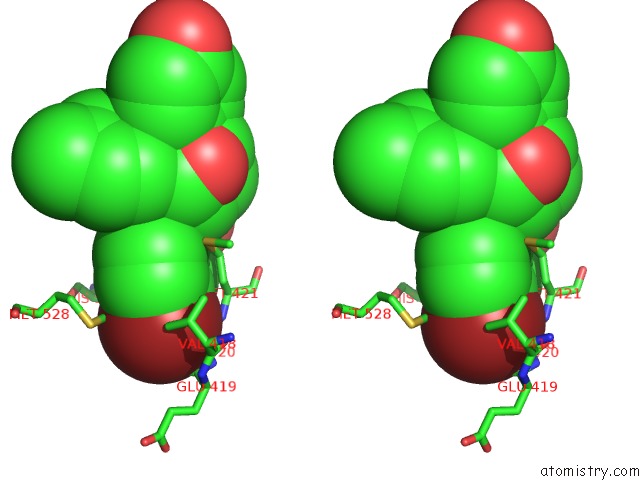
Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 1 of Crystal Structure of the Er-Alpha Ligand-Binding Domain (Y537S) in Complex with the Obhs-Asc Analog, (E)-6-(4-((1R,4S,6R)-6-((4- Bromophenoxy)Sulfonyl)-3-(4-Hydroxyphenyl)-7-Oxabicyclo[2.2.1]Hept-2- En-2-Yl)Phenyl)Hex-5-Enoic Acid within 5.0Å range:
|
Reference:
J.C.Nwachukwu,
S.Srinivasan,
N.E.Bruno,
J.Nowak,
N.J.Wright,
F.Minutolo,
E.S.Rangarajan,
T.Izard,
X.Q.Yao,
B.J.Grant,
D.J.Kojetin,
O.Elemento,
J.A.Katzenellenbogen,
K.W.Nettles.
Systems Structural Biology Analysis of Ligand Effects on Er Alpha Predicts Cellular Response to Environmental Estrogens and Anti-Hormone Therapies. Cell Chem Biol V. 24 35 2017.
ISSN: ESSN 2451-9456
PubMed: 28042045
DOI: 10.1016/J.CHEMBIOL.2016.11.014
Page generated: Mon Jul 7 09:16:57 2025
ISSN: ESSN 2451-9456
PubMed: 28042045
DOI: 10.1016/J.CHEMBIOL.2016.11.014
Last articles
Ca in 1AVWCa in 1AVC
Ca in 1AVH
Ca in 1AUI
Ca in 1AUX
Ca in 1AUJ
Ca in 1ATJ
Ca in 1ATN
Ca in 1AU9
Ca in 1ATL