Bromine »
PDB 5v2h-5y93 »
5vij »
Bromine in PDB 5vij: Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc
Protein crystallography data
The structure of Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc, PDB code: 5vij
was solved by
T.-C.Mou,
P.Conti,
A.Pinto,
L.Tamborini,
S.R.Sprang,
K.B.Hansen,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 19.98 / 2.11 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 54.886, 87.158, 122.254, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.3 / 25.5 |
Bromine Binding Sites:
The binding sites of Bromine atom in the Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc
(pdb code 5vij). This binding sites where shown within
5.0 Angstroms radius around Bromine atom.
In total only one binding site of Bromine was determined in the Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc, PDB code: 5vij:
In total only one binding site of Bromine was determined in the Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc, PDB code: 5vij:
Bromine binding site 1 out of 1 in 5vij
Go back to
Bromine binding site 1 out
of 1 in the Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc
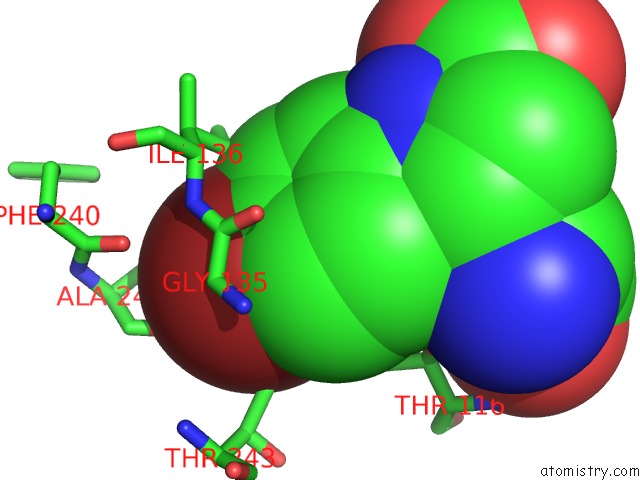
Mono view
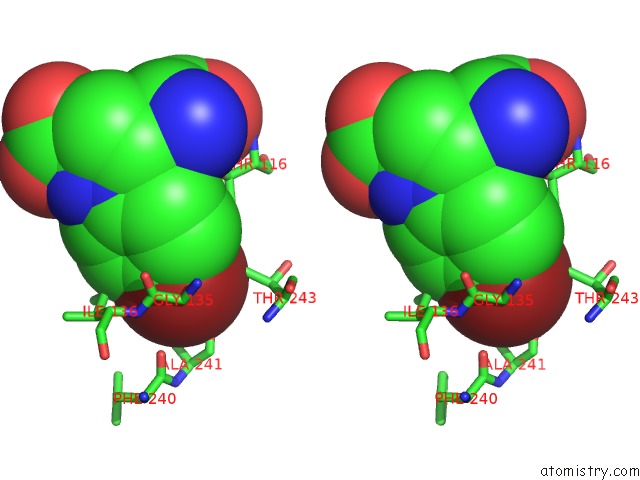
Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 1 of Crystal Structure of GLUN1/GLUN2A Nmda Receptor Agonist Binding Domains with Glycine and Antagonist, 4-Bromophenyl-Acepc within 5.0Å range:
|
Reference:
G.E.Lind,
T.C.Mou,
L.Tamborini,
M.G.Pomper,
C.De Micheli,
P.Conti,
A.Pinto,
K.B.Hansen.
Structural Basis of Subunit Selectivity For Competitive Nmda Receptor Antagonists with Preference For GLUN2A Over GLUN2B Subunits. Proc. Natl. Acad. Sci. V. 114 E6942 2017U.S.A..
ISSN: ESSN 1091-6490
PubMed: 28760974
DOI: 10.1073/PNAS.1707752114
Page generated: Mon Jul 7 09:21:47 2025
ISSN: ESSN 1091-6490
PubMed: 28760974
DOI: 10.1073/PNAS.1707752114
Last articles
Cl in 5RSLCl in 5ROW
Cl in 5RMJ
Cl in 5ROP
Cl in 5RML
Cl in 5RKR
Cl in 5RKE
Cl in 5RH4
Cl in 5RH3
Cl in 5RH2