Bromine »
PDB 8qs8-8u3a »
8tsb »
Bromine in PDB 8tsb: Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2
Protein crystallography data
The structure of Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2, PDB code: 8tsb
was solved by
M.Holliday,
Y.Tang,
A.Bulku,
J.Wilbur,
J.Fraser,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 49.54 / 3.53 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 87.871, 119.947, 189.317, 90, 90, 90 |
| R / Rfree (%) | 21.6 / 25.6 |
Other elements in 8tsb:
The structure of Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2 also contains other interesting chemical elements:
| Fluorine | (F) | 1 atom |
Bromine Binding Sites:
The binding sites of Bromine atom in the Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2
(pdb code 8tsb). This binding sites where shown within
5.0 Angstroms radius around Bromine atom.
In total only one binding site of Bromine was determined in the Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2, PDB code: 8tsb:
In total only one binding site of Bromine was determined in the Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2, PDB code: 8tsb:
Bromine binding site 1 out of 1 in 8tsb
Go back to
Bromine binding site 1 out
of 1 in the Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2

Mono view
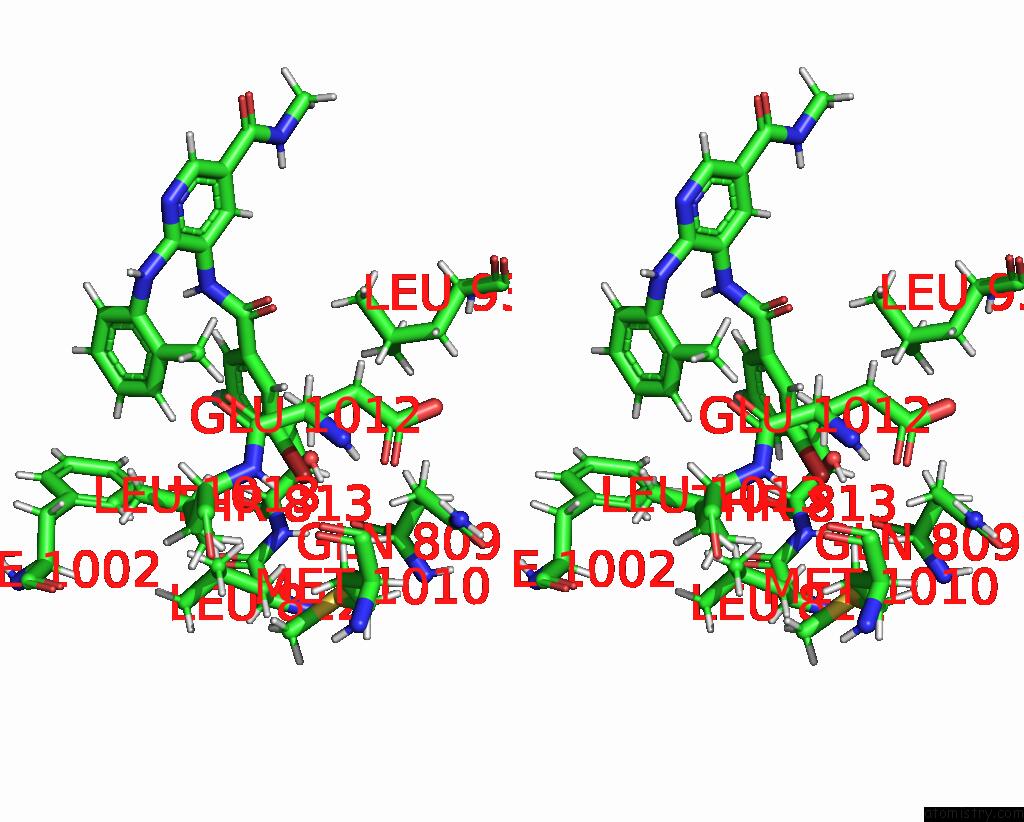
Stereo pair view

Mono view

Stereo pair view
A full contact list of Bromine with other atoms in the Br binding
site number 1 of Human PI3K P85ALPHA/P110ALPHA Bound to Compound 2 within 5.0Å range:
|
Reference:
A.Varkaris,
E.Pazolli,
H.Gunaydin,
Q.Wang,
L.Pierce,
A.A.Boezio,
L.Dipietro,
A.Frost,
F.Giordanetto,
E.P.Hamilton,
K.Harris,
M.Holliday,
T.L.Hunter,
A.Iskandar,
Y.Ji,
A.Larivee,
J.R.Larochelle,
A.Lescarbeau,
F.Llambi,
B.Lormil,
M.M.Mader,
B.G.Mar,
I.Martin,
T.H.Mclean,
K.Michelsen,
Y.Pechersky,
E.Puente-Poushnejad,
R.Samadani,
A.M.Schram,
K.Shortsleeves,
S.Swaminathan,
S.Tajmir,
G.Tan,
Y.Tang,
R.Valverde,
B.Wehrenberg,
J.Wilbur,
B.R.Williams,
H.Zeng,
W.P.Walters,
B.B.Wolf,
D.E.Shaw,
D.A.Bergstrom,
J.Watters,
J.S.Fraser,
P.D.Fortin,
D.R.Kipp.
Discovery and Clinical Proof-of-Concept of Rly-2608, A First-in-Class Mutant-Selective Allosteric PI3KA Inhibitor That Decouples Anti-Tumor Activity From Hyperinsulinemia. Cancer Discov 2023.
ISSN: ESSN 2159-8290
PubMed: 37916956
DOI: 10.1158/2159-8290.CD-23-0944
Page generated: Mon Jul 7 12:38:26 2025
ISSN: ESSN 2159-8290
PubMed: 37916956
DOI: 10.1158/2159-8290.CD-23-0944
Last articles
F in 7GORF in 7GOV
F in 7GON
F in 7GOB
F in 7GO1
F in 7GO7
F in 7GN2
F in 7GMZ
F in 7GM4
F in 7GMY